This is part of my “journal club for credit” series. You can see the other computational neuroscience papers in this post.
Unit: Deep Learning
- Perceptron
- Energy Based Neural Networks
- Training Networks
- Deep Learning
Paper
The Perceptron: A Probabilistic Model for Information Storage and Organization in the Brain by Rosenblatt in 1958
Other Useful References
Motivation for Perceptron
I’ll let Rosenblatt introduce the important questions leading to the perceptron himself by quoting his first paragraph:
If we are eventually to understand the capability of higher organisms for perceptual recognition, generalization, recall, and thinking, we must first have answers to three fundamental questions:
- How is information about the physical world sensed, or detected, by biological system?
- In what form is information stored, or remembered?
- How does information in storage, or in memory, influence recognition and behavior?
The perceptron is a first attempt to answer second and third questions. In the years leading up to the perceptron, there were two dominate themes of theoretical research on the brain. One focused on the general computational properties of the brain (McCollough and Pitts 1943) and showed that simple binary neurons could form a computer (ie they can compute any possible function). Another theme focused on abstracting away the details of experiments to get at general principles that relate to computation in the brain (Hebb 1949 and his synapse learning rules).
The perceptron opened up a third avenue of theoretical research. The central goal is to devise neuron-inspired algorithms that learn from real data and can be used to make a decision.
What is a Perceptron?
Basics
I find the math in the original perceptron paper pretty confusing. This is partly due to a generational difference in terminology, and partly due to poor explanations in the paper. This is definitely a paper that benefited from the passage of time and future synthesis into a more concise topic. Therefore, I recommend focusing attention on the introduction and conclusion, while below I’ll introduce the modern notation of the perceptron (see MacKay Ch 39/40 for similar details).

The perceptron consists of a set of inputs,  , that are fed into the perceptron, with each input receiving its own weight,
, that are fed into the perceptron, with each input receiving its own weight, 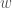 . The activity of the percepton is given by
. The activity of the percepton is given by 
Note that the perceptron can have a bias that is independent of inputs. However, we don’t need to write this out separately and can instead include an input that is always set to 1, independently of the data.
This activity is then evaluated by the activation function,  , to determine the output,
, to determine the output, 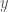 . There are lots of different possible activation rules with some popular ones including
. There are lots of different possible activation rules with some popular ones including




The end result is that we can take the output of a perceptron and use this output to make a decision. The sigmoid and threshold activation functions return an answer between 0 and 1 and hence have a natural interpretation as a probability. From now on, we will work with sigmoid activation functions.
Training
Now that the basics of a perceptron have been introduced, how do we actually train it? In other words, if I gave you a set of data, 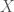 , where each entry
, where each entry 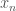 is
is 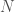 dimensional, how would I evaluate perceptron’s handling of the data? For now, we will focus on using the perceptron as a binary classifier (only need to decide between two groups: 0 and 1). Since we are using sigmoid activation functions, we can interpret the output as the probability that the data is in group 1.
dimensional, how would I evaluate perceptron’s handling of the data? For now, we will focus on using the perceptron as a binary classifier (only need to decide between two groups: 0 and 1). Since we are using sigmoid activation functions, we can interpret the output as the probability that the data is in group 1.
The standard way to train a binary classier is to have a training set which consists of pairs of data, 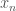 , and correct labels,
, and correct labels, 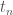 . Then training proceeds by seeing if the output label of a perceptron matches the correct label. If everything is correct, perfect! If not, we need to do something with that error.
. Then training proceeds by seeing if the output label of a perceptron matches the correct label. If everything is correct, perfect! If not, we need to do something with that error.
For a sigmoid activation, the commonly used error function is the cross-entropy:
![\mathcal{E} = - \sum_n \left[ t_n \ln y_n + \left(1-t_n \right) \ln \left(1-y_n \right) \right]](https://s0.wp.com/latex.php?latex=%5Cmathcal%7BE%7D+%3D+-+%5Csum_n+%5Cleft%5B+t_n+%5Cln+y_n+%2B+%5Cleft%281-t_n+%5Cright%29+%5Cln+%5Cleft%281-y_n+%5Cright%29+%5Cright%5D+&bg=ffffff&fg=444444&s=0&c=20201002)
The output 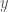 is a function of the weights
is a function of the weights 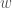 . We can then take the derivate of the error with respect to the weights, which in the case of the sigmoid activation and cross-entropy error is simply
. We can then take the derivate of the error with respect to the weights, which in the case of the sigmoid activation and cross-entropy error is simply 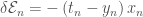 .
.
The simplest possible update algorithm is to perform gradient descent on the weights and define  . This is a greedy algorithm (always improves current error, longterm consequences be damned!). Gradient descent comes in several closely related varieties: online, batch, and mini-batch. Let’s start with the mini-batch. First the data is divided up into small random sets (say 10 data points each). Then we loop through the mini-batches, and for each one we calculate the output and error and update the weights. Online learning is when the mini-batches each contain exactly 1 data point, while batch learning is when the mini-batch is the whole dataset. The current standard is to use a mini-batch of between 10 to 100 which is a compromise between speed (batches are faster) and accuracy (online finds better solutions).
. This is a greedy algorithm (always improves current error, longterm consequences be damned!). Gradient descent comes in several closely related varieties: online, batch, and mini-batch. Let’s start with the mini-batch. First the data is divided up into small random sets (say 10 data points each). Then we loop through the mini-batches, and for each one we calculate the output and error and update the weights. Online learning is when the mini-batches each contain exactly 1 data point, while batch learning is when the mini-batch is the whole dataset. The current standard is to use a mini-batch of between 10 to 100 which is a compromise between speed (batches are faster) and accuracy (online finds better solutions).
Putting it all together, the training algorithm is as follows
- Calculate the activation function and output with respect to a mini-batch of data
- Calculate the errors of the output
- Update the weights
And now you’ve got all the basics of a perceptron down! On to the more difficult questions…
Fundamental Questions
- What are similarities and differences between a perceptron and a neuron? Do different activation functions lead to distinct interpretations?
- What is connectivisim? How does this relate to the perceptron? How does this contrast with computers?
- What class of learning algorithm is the perceptron? Possible answers: unsupervised, supervised, or reinforcement learning
- What type of functions can a perceptron compute? Compare the standard OR gate vs the exclusive OR (XOR) gate for a perceptron with 2 weights.
- Does the perceptron return a unique answer? Does the perceptron return the “best” answer (you need to define “best”)? Check out Support Vector Machines for one answer to the “best”.
- Under what conditions can the perceptron generalize to data it has never seen before? Look into Rosenblatt’s “differentiated environment”.
Additional Questions
- There are other possibilities for the error functions. Why is the cross-entropy a wise choice for the sigmoid activation?
- The weight updates can be multiplied by a “learning rate” that controls the size of updates, while I implicitly assumed a learning rate of 1. How would you actually determine a sensible learning rate?
- The standard learning algorithm puts no constraints on the possible weights. What are some possible problems with unconstrained weights? Can you think of a possible solution? How does this change the generalization properties of a perceptron?
- Threshold activation functions produce simpler output (only two possible values) than sigmoid activation functions. Despite this simpler output, threshold activation functions are more difficult to train. Can you figure out why?
- What is the information storage capacity of a perceptron? The exact answer is difficult, but you can get the right order of magnitude in the limit of large number of data points and large number of weights.
